Nuclear Hormone Receptors: A Brief Overview
Understand Nuclear Receptors through our introduction to this family of intracellular transcription factors that regulate gene expression.
Introduction

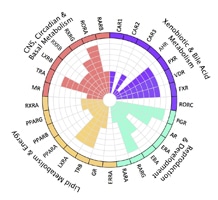
Figure 1. Phylogenetic classification of nuclear receptors
Unlike receptors found on the cell surface, members of the nuclear hormone receptor (NR) superfamily are restricted to metazoan organisms such as nematodes, insects, and vertebrates. These proteins are intracellular transcription factors that directly regulate gene expression in response to lipophilic molecules. They affect a wide variety of functions, including fatty acid metabolism, reproductive development, and detoxification of foreign substances. Many of the NRs act as ligand-inducible transcription factors, responding to endogenous and exogenous chemicals. However, the vast majority of known receptors do not have an identifiable physiologically relevant ligand, and are deemed orphan receptors. To date, over 300 NRs have been cloned, although there are only 48 members of the family in humans. Early classification of these receptors was based on ligands, DNA binding properties or other functional characterization. A more systematic classification has been proposed, based on sequence similarity, much like that employed for cytochrome P450s. Phylogenic analysis has shown six subfamilies (NR1-6) with various groups and individual genes. (1) (See Figure 1). As discussed below, most NRs have the same basic structure. The most highly conserved region is the C4 zinc finger domain, which as described above, is a DNA binding motif.
Structure and Function
Functional domains
NRs generally follow a standard blueprint as shown in Figure 2 (reviewed in 2-4). While most NRs have all of these elements, there are a number of exceptions, some of which will be discussed below. The N terminus of the NR, sometimes called the modulator, hypervariable or A/B domain, has transactivation activity, termed activation function 1 (AF-1). This acidic activation domain (AD) is ligand-independent, or constitutively functional. The A/B domain’s sequence and length are highly variable between receptors (i.e., GR versus RXR) and among receptor subtypes (RXRα versus RXRβ). In addition, this region is the most frequent site of alternative splicing and secondary start sites and contains a variety of kinase recognition sequences. For these reasons, it is thought that the variable N- terminal sequences may be responsible for the receptor-, species-, and cell type-specific effects as well as promoter context-dependent properties of NR transactivation.
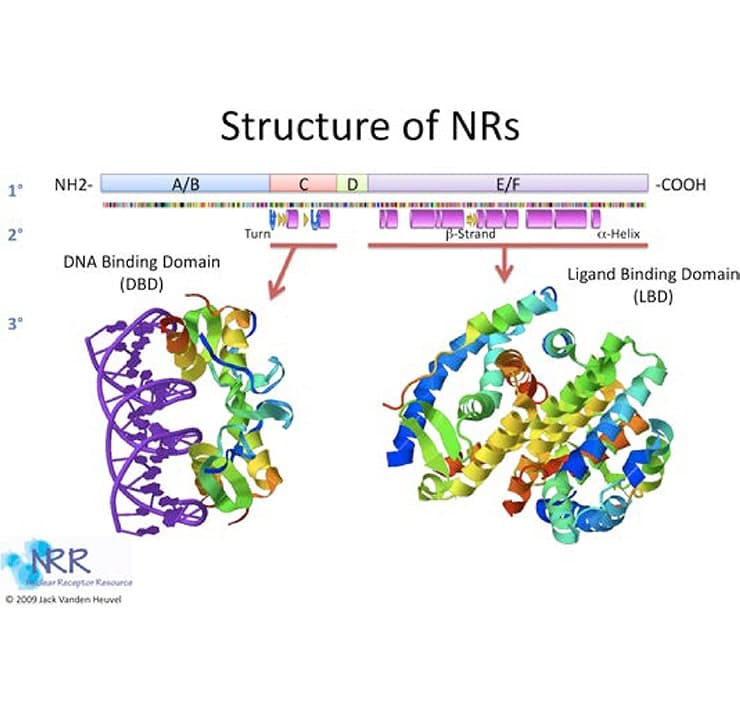
Figure 2. Primary, secondary and tertiary structure of nuclear receptors
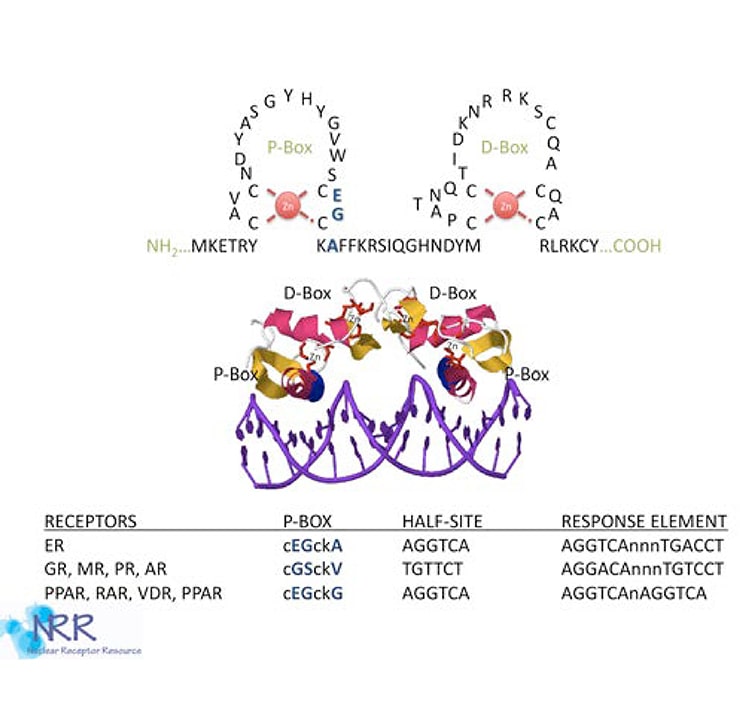
Figure 3. The NR DNA binding domain
NRs bind to hormone response elements (HREs) in their target promoters through the DNA binding domain (DBD) or C domain. Composed of two zinc fingers, the DBD is the most conserved region within the NR superfamily (see Figure 3). The first zinc finger contains the proximal-or P-box region, an alpha helix that is responsible for high-affinity recognition of the “core half-site” of the response element. Located within the second zinc finger is the distal or D-box, an α-helix which lies perpendicular to P-box helix, and is a site that mediates receptor dimerization. NRs bind to DNA as heterodimers, homodimers, or monomers, depending on the class of NR. The steroid hormone receptors GR, PR, ER, AR and MR (receptors for glucocorticoid, progesterone, estrogen, androgen, and mineralocorticoids, respectively) bind to DNA as homodimers and recognize palindromic response elements. However, thyroid, retinoid, vitamin D, and peroxisome proliferator receptors (TR, RAR, VDR and PPAR), as well as most orphan receptors, bind to DNA as a heterodimer with retinoid-x-receptor (RXR). However, the three dimensional structure of the RXR heterodimer complex produces different DNA binding affinities. Response elements may be direct repeats (DRx, AGGTCA-Nx-AGGTCA, where N is any nucleotide and x is any number of residues from 0-10), everted repeats (ERx, ACTGGA-Nx-AGGTCA) or inverted repeat (IRx, AGGTCA-Nx-ACTGGA).
Immediately adjacent to the DNA binding domain is the D or hinge domain. This particular region has an ill-defined function. The hinge domain contains the carboxy-terminal extension (CTE) of the DBD, which may be involved in recognizing the extended 5’ end of the HRE. The D-domain appears to allow for conformational changes in the protein structure following ligand binding. Also, this region may contain nuclear localization signals and protein-protein interaction sites.
The sequence of the ligand binding domain (LBD) or E/F domain varies substantially between NRs, but they all share a common structure of 11-13 a-helices organized around a hydrophobic binding pocket. Residues within the binding pocket confer specificity, determining whether the LBD will accept steroid hormones, retinoid compounds, or the host of xenobiotic ligands that affect receptor function. Ligand-dependent activation requires the presence of activation function 2 (AF-2), located at the extreme C terminus of the NR. LBDs also contain nuclear localization signals, protein interaction with dimerization motifs for heat shock proteins, coregulators and other transcription factors.
Exceptions to the Rule
There are several NRs that do not follow the basic structure function paradigm. For example, the constitutively active receptor (CAR) allows constitutive transcription of the target gene in the absence of ligand. Transcriptional activity is suppressed by binding to ligands androstenol and androstanol, unlike other NRs examined. Another important exception to the classic model of NR activation comes from receptors that do not require DNA binding. DAX-1 and SHP lack DBDs completely, and transgenic mice bearing a GR that does not bind DNA are viable and fertile even though at least some GR functions are crucial for survival. NRs can also have important biological effects without ligand binding. Most NRs are phosphoproteins, and it is now known that many of these proteins are activated by crosstalk with other signal transduction pathways such as those responding to EGF and TGF-α.
Basic Mechanism of Action
The mechanism of action of nuclear hormone receptors can take one of two basic forms, that of steroid hormone receptors (SHRs) or that of retinoid/thyroid/Vitamin D receptors (see Figure 4, reviewed in (5-7). In the absence of ligand, the transcriptionally inactive SHRs MR, PR, GR, AR, and ER are sequestered in a large complex comprising the receptor, heat shock protein-90 (HSP90), Hsp70, FKBP52/51 and possibly other proteins. The cellular localization of this inactive complex is somewhat controversial and cytoplasmic or nuclear localization may be observed depending on the cell type and the conditions examined; however, the central dogma is that SHRs are cytosolic in the un-liganded form. One consequence of hormone binding to the receptor is a distinct conformational change in receptor structure (discussed below). This conformational change marks the beginning of the signal transduction process. In the case of the GR subfamily (GR, AR, MR, PR), hormone binding elicits a dissociation of hsps and the release of a monomeric receptor from the complex. Genetic analysis and in vitro protease digestion experiments indicate that the conformational changes in receptor structure induced by agonists are similar but distinct from those produced by antagonists. However, both conformations appear to be incompatible with hsp binding.
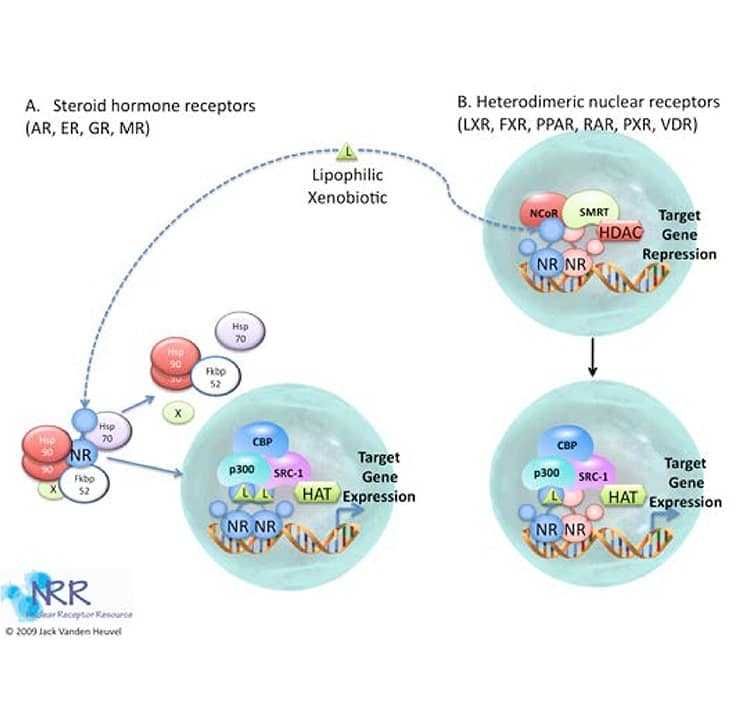
Figure 4. Basic mechanism of action of nuclear receptors
The TR, RAR, and VDR receptors do not avidly interact with hsps and are localized predominantly in the nucleus in the absence of ligand. Some unliganded NRs of this class may interact with DNA and act as transcription repressors. This may be the result of interaction with co-repressor proteins. An interesting exception to this observation is the constitutively active receptor (CAR) that is transcriptionally active in the absence of its ligand. Hormone induced conformational changes also occur upon activation of this class of NR, suggesting that alteration of receptor shape by ligands is a key step in the activation pathway.
Evidence suggests that receptors of the GR subfamily (SHRs) cooperatively bind to DNA as homodimers. The TR, RAR, VDR, PPAR and most of the orphan receptors form heterodimers with other members of the intracellular receptor superfamily. TR, RAR, PPAR and VDR can utilize RXRs as partners for heterodimer formation. The DNA site of contact depends on certain sequences within the C-domain, namely the proximal (P-box) and distal (D-box) zinc finger motifs (see description of the C-domain above). The P-box determines the half-site recognized, while the D-box determines the spacing between half-sites. Following activation, the SHRs receptors are capable of interacting with DNA, and both classes of NRs (SHRs and TR/RAR) can now recruit co-activators. The DNA bound NR complex is now a substrate for general transcription apparatus and the initiation of transcription commence.
Ligands and Activators
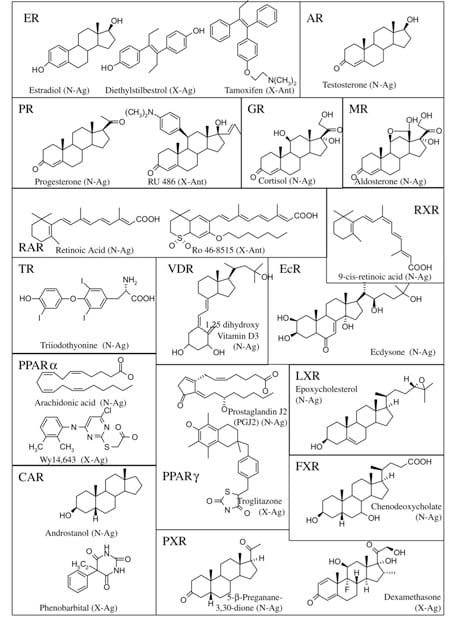
Figure 5. Representative NR ligands
Natural and xenobiotic ligands
Ligands for NRs are as varied as the proteins themselves (Figure 5). Although the structures of these compounds is varied, a few generalized comments can be made. All ligands are lipophilic and can easily transverse the plasma membrane as well as the nuclear membrane, if required. The affinity (Kd) of the ligand-receptor complex is generally in the nM range, but can vary from pM to mM. It should be noted however, that the concentrations of each natural ligand should approach their Kd to be considered a physiologically-relevant ligand. This is of particular importance when considering reclassifying (or adopting) an orphan receptor in the process of reverse endocrinology. Xenobiotic agonists and antagonists share structure features with the natural effector molecules, although the similarities are often cryptic. Some receptors, such as PPARγ, have a large ligand-binding cavity that allows for the association of a variety of endogenous ligands.
Ligand-induced activation
Structural studies of empty and ligand-bound LBDs have led to the “mousetrap” model of NR activation (Figure 6). The ligand is attracted to the trap, the receptor’s electrostatic potential, and a conformational change takes place, preventing the ligand’s exit. In the same way that the sprung mousetrap is more stable than the primed trap, ligand binding to the NRs ligand-binding domains stabilizes their structures relative to the unliganded receptor. The ligand forms an integral part of the hydrophobic core of the liganded LBD. This structural change is different for ligands that are full agonists versus those that are partial agonists or antagonists. Much attention is focused on the accessibility of the AF-2 domain to accessory proteins. The AF-2 domain can serve as an activator of transcription when excised from the rest of the protein and linked to a heterologous DNA-binding domain. In this model, binding of the ligand molecule induces a conformational change in the LBD, whereby the AF-2 sequences fold back against the binding pocket, obstructing the opening and causing rearrangements in adjacent helices. In the process, a new surface is revealed that recruits specific transcriptional coactivators. This model may explain why receptor antagonists block transactivation; these compounds do not induce the proper conformational rearrangements in the LBD, interfering with the formation of the transcriptional activation complex.
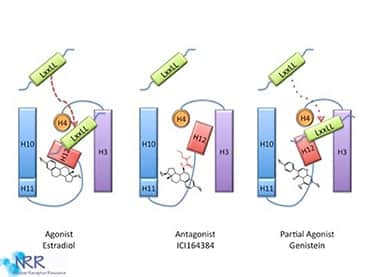
Figure 6. Ligand binding to NRs results in a conformational change in the protein
Reverse Endocrinology
The increasing use of bioinformatics and the spread of genome projects has led to the discovery of hundreds of proteins which share structural characteristics with NRs. When a NR is discovered without any knowledge of its natural ligand, it is dubbed an orphan nuclear receptor (ONR). Efforts to understand ONR function and identify their physiological ligands (a process known as reverse endocrinology) have led to the discovery of novel metabolic pathways involving the PPAR, liver X receptor (LXR), and farnesoid X receptor (FXR), novel classes of ligands (benzoates, terpenoids), and alternative mechanisms for NR receptor regulation and function.
The cloning of the intracellular receptor cDNAs and their reconstruction into ligand responsive transcription units has allowed for the identification of new agonist/antagonists. The “cis-trans” or reporter assay consists of transfecting a receptor expression vector and a vector carrying a receptor-responsive reporter into mammalian cells. Induction of the reporter gene activity (i.e. luciferase, chloramphenicol acetyltransferase, b-galactosidase) reflects the transcriptional activity of the transfected receptor induced by hormone. The identification of receptor antagonists is more complex as it is sometimes difficult to distinguish true receptor antagonism from toxicity. The domain structure of NRs allows for construction of reporter assays without knowledge of the DNA binding domain of the protein. A common means to assess ligand activation of a receptor is to perform a “finger-swap” whereby the LBD for the novel NR is placed downstream of a heterologous DBD of another, well-characterized protein. Examples of DBD used include the C-domain of GR as well as the transcription factor Gal4. Activation of this chimeric protein by agonists for the novel LBD would regulate a reporter that contains the appropriate DNA binding site (i.e., a GRE for proteins containing the C-domain of GR). This type of approach allows for rapid screening of chemicals and may be used to find both xenobiotic and endogenous ligands.
Ligand-independent Activation
An emerging concept for intracellular receptors is that they function not only as transducers of nuclear effects of steroids, hormones and nutrients, but also as key points of convergence of multiple signal transduction pathways (reviewed in (8-12)). The first realization of this “cross-talk” between signaling cascades came from the observation that transcriptional activity of PR, ER, TR, and COUP-TF is modulated by the neurotransmitter dopamine. The specific activation of the dopamine D1 receptors results in activation of at least two distinct signal transduction processes in the cell. Mutation of a specific serine site close to the carboxyl terminus of the NR can prevent dopamine activation, with no effect on the ability of the ligand to activate. The two activation pathways may differentially phosphorylate this serine. A similar phenomenon is seen with other signaling pathways, in particular polypeptide growth factors such as insulin-like growth factor (IGF-1), transforming growth factor a (TGFa) and epidermal growth factor (EGF).
Citations
- Nuclear Receptors Nomenclature Committee (1999) A unified nomenclature system for the nuclear receptor superfamily, Cell 97, 161-163.
- Chawla, A., Repa, J. J., Evans, R. M., and Mangelsdorf, D. J. (2001) Nuclear receptors and lipid physiology: opening the X-files, Science 294, 1866-1870.
- Bourguet, W., Germain, P., and Gronemeyer, H. (2000) Nuclear receptor ligand-binding domains: three-dimensional structures, molecular interactions and pharmacological implications, Trends Pharmacol Sci 21, 381-388.
- Wahli, W., and Martinez, E. (1991) Superfamily of steroid nuclear receptors: positive and negative regulators of gene expression, Faseb J 5, 2243-2249.
- Francis, G. A., Fayard, E., Picard, F., and Auwerx, J. (2003) Nuclear receptors and the control of metabolism, Annu Rev Physiol 65, 261-311.
- Khan, S. A., and Vanden Heuvel, J. P. (2003) Role of nuclear receptors in the regulation of gene expression by dietary fatty acids (review), J Nutr Biochem 14, 554-567.
- Smirnov, A. N. (2002) Nuclear receptors: nomenclature, ligands, mechanisms of their effects on gene expression, Biochemistry (Mosc) 67, 957-977.
- Ward, R. D., and Weigel, N. L. (2009) Steroid receptor phosphorylation: Assigning function to site-specific phosphorylation, Biofactors.
- Weigel, N. L., and Moore, N. L. (2007) Steroid receptor phosphorylation: a key modulator of multiple receptor functions, Mol Endocrinol 21, 2311-2319.
- Weigel, N. L., and Moore, N. L. (2007) Kinases and protein phosphorylation as regulators of steroid hormone action, Nucl Recept Signal 5, e005.
- Weigel, N. L., and Zhang, Y. (1998) Ligand-independent activation of steroid hormone receptors, J Mol Med 76, 469-479.
- Weigel, N. L. (1996) Steroid hormone receptors and their regulation by phosphorylation, Biochem J 319 ( Pt 3), 657-667.