Transrepression Assays in Drug Discovery and Mechanistic Pharmacology
Transrepression assays are specialized cell-based systems designed to measure functional suppression of gene expression mediated by nuclear receptors and transcription factors. While traditional reporter assays quantify transcriptional activation, transrepression assays are used to evaluate how a receptor or pathway reduces transcription factor-driven gene expression. These assays are especially relevant in autoimmune diseases, inflammation research, immunology, and safety assessment, where gene suppression rather than activation often drives therapeutic benefit.

Transrepression typically occurs through protein–protein interactions rather than direct DNA binding. In many cases, a ligand-activated nuclear receptor interferes with pro-inflammatory transcription factors, preventing them from initiating gene transcription. A well-characterized example involves the Glucocorticoid Receptor. Upon ligand binding, this receptor can repress the activity of transcription factors such as NF-κB and AP-1, reducing expression of cytokines and other inflammatory mediators. Importantly, this repression often occurs independently of direct glucocorticoid response element binding, instead relying on transcription factor cross-talk and cofactor competition.
Mechanistic Basis of Transrepression
At the molecular level, transrepression may involve several non-mutually exclusive mechanisms. Ligand-bound nuclear receptors can interfere with pro-inflammatory transcription factors, block recruitment of coactivators, promote recruitment of corepressors, or interfere with chromatin remodeling complexes. In some contexts, receptors also modulate signaling cascades upstream of transcription factor activation, indirectly dampening transcriptional output.
Cell-based transrepression assays are commonly built using luciferase or comparable reporter genes controlled by transcription factor–responsive promoter elements. Cells are stimulated with a defined activator, such as a cytokine or Toll-like receptor agonist, to induce transcription factor signaling. Test compounds are then added to determine whether they reduce the induced signal. A decrease in reporter output relative to the stimulated control is interpreted as functional repression, provided that parallel controls rule out cytotoxicity or nonspecific suppression of reporter expression. Because these assays operate in intact cells, they can capture integrated effects of receptor engagement, signaling dynamics, cofactor usage, and transcriptional cross-talk.
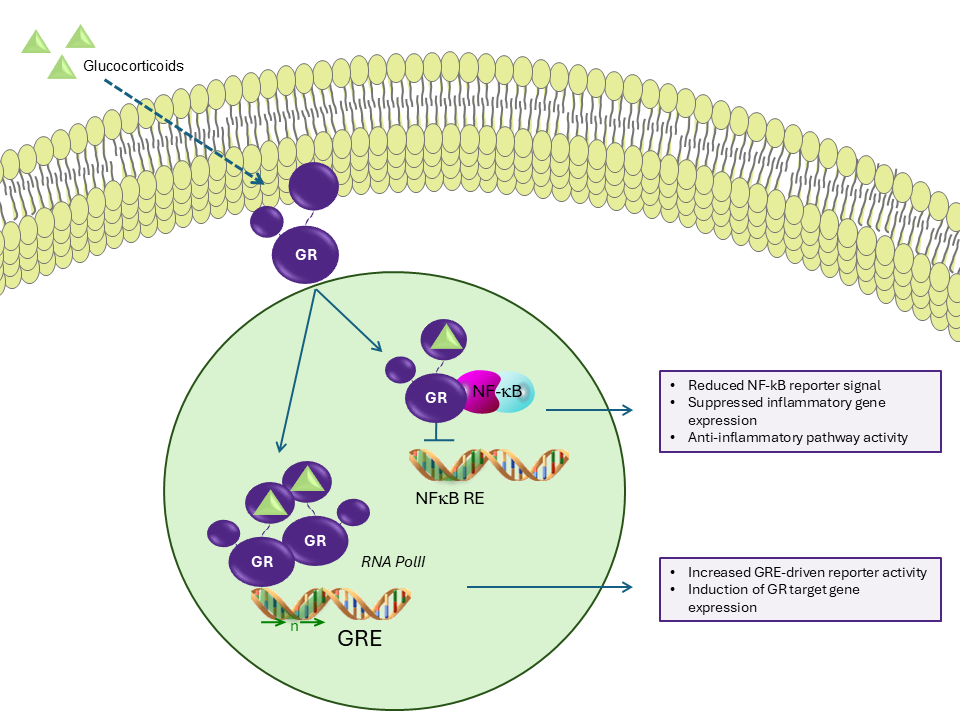
Distinguishing Transactivation from Transrepression
In nuclear receptor pharmacology, it is useful to differentiate between transactivation and transrepression. Transactivation assays measure receptor-driven induction of gene expression through response elements or activation-associated regulatory regions, whereas transrepression assays measure reduction of transcription factor-driven reporter activity relative to an activated baseline.
This distinction has been particularly influential in glucocorticoid receptor drug discovery, where selective modulators have historically been evaluated for the ability to retain anti-inflammatory repression while reducing liabilities associated with broader transcriptional activation programs. This concept also underpins interest in Selective Glucocorticoid Receptor Modulators (SGRMs), which aim to preserve anti-inflammatory transrepression while reducing transactivation-associated side effects. Functional assays capable of independently quantifying these two activities are therefore central to biased agonism studies and structure–activity relationship optimization.
Applications in Inflammation and Immunology
Transrepression assays are widely used to study inflammatory signaling pathways. Suppression of NF-κB and AP-1 activity is relevant across autoimmune disease, respiratory inflammation, dermatological conditions, and metabolic disorders. Because these assays measure upstream transcriptional regulation rather than downstream cytokine secretion, they provide mechanistic resolution that complements protein-level measurements such as ELISA. This upstream positioning can be particularly valuable during early-stage screening, where distinguishing primary pathway modulation from secondary effects is critical.
Nuclear Receptor Cross-Talk and Selective Modulation
Multiple nuclear receptors beyond the Glucocorticoid Receptor (GR) exhibit transrepressive behavior. The Peroxisome Proliferator-activated Receptor Gamma (PPARγ) has been shown to modulate inflammatory gene expression through interference with pro-inflammatory transcription factors. Similarly, Estrogen Receptor Alpha (ERα) and Androgen Receptor (AR) can influence immune signaling pathways in certain cellular contexts. Transrepression assays allow investigators to quantify these effects directly and evaluate ligand-dependent differences in pathway modulation.
Such functional assays are particularly helpful during lead optimization. Compounds that exhibit comparable binding affinity may produce different repression profiles depending on conformational effects, coregulator recruitment, or cell-type–specific signaling context. Quantitative transrepression data therefore contribute to mechanistic clarity beyond receptor binding or simple potency measurements alone.
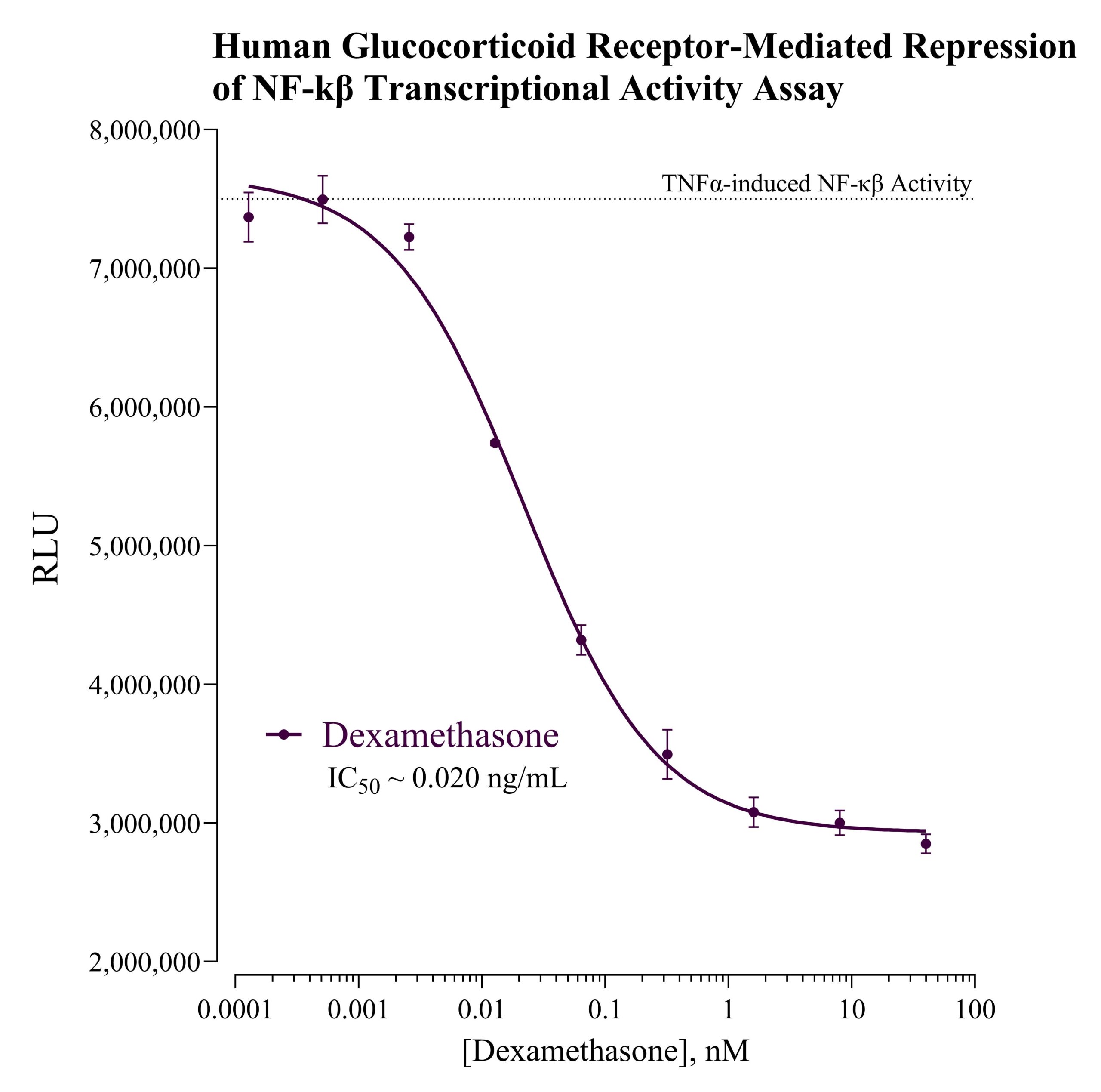
Role in Toxicology and Safety Assessment
In mechanistic toxicology, transcription factor repression assays provide insight into how xenobiotics alter inflammatory or endocrine signaling networks. Because the assays operate in intact cells and measure integrated pathway responses, they can reveal subtle modulatory effects that may not be apparent from receptor-binding or enzymatic assays alone. For this reason, they can support hypothesis generation around off-target activity, immunomodulatory effects, and pathway-selective liabilities.
Experimental Considerations
Successful implementation of transrepressor assays requires careful control of stimulus intensity, timing, and cellular context. Overstimulation may mask partial repression, whereas insufficient activation reduces dynamic range. Selection of appropriate cell lines, reporter constructs, and assay windows is critical for reproducibility. Stable reporter cell lines often enhance consistency across experiments, particularly in longitudinal studies or high-throughput screening environments.
Data analysis typically involves normalization to stimulated controls and calculation of concentration–response parameters such as IC₅₀ values. Interpretation should account for potential cytotoxicity or nonspecific effects that could artificially reduce reporter signal. Parallel viability measurements are therefore recommended.
Transrepression assays provide a useful functional platform for quantifying transcriptional suppression in living cells. Their strength lies in measuring integrated pathway behavior rather than isolated molecular events. When designed and interpreted carefully, these assays can support mechanistic studies in inflammation, nuclear receptor pharmacology, immunology, and toxicology, and they remain valuable tools for evaluating pathway-selective modulation during drug discovery.
References
Gerber, A. N., Newton, R., & Sasse, S. K. (2021). Repression of transcription by the glucocorticoid receptor: A parsimonious model for the genomics era. Journal of Biological Chemistry, 296, 100687. https://doi.org/10.1016/j.jbc.2021.100687
Hudson, W. H., de Vera, I. M. S., Nwachukwu, J. C., Weikum, E. R., Herbst, A. G., Yang, Q., Bain, D. L., Nettles, K. W., Kojetin, D. J., & Ortlund, E. A. (2018). Cryptic glucocorticoid receptor-binding sites pervade genomic NF-κB response elements. Nature Communications, 9, 1337. https://doi.org/10.1038/s41467-018-03780-1
Jakob, F., Hennen, S., Gautrois, M., Khalil, F., & Lockhart, A. (2025). Novel selective glucocorticoid receptor modulator GRM-01 demonstrates dissociation of anti-inflammatory effects from adverse effects on glucose and bone metabolism. Frontiers in Pharmacology, 16. https://doi.org/10.3389/fphar.2025.1542351